-Search query
-Search result
Showing 1 - 50 of 130 items for (author: huber & st)

EMDB-17863: 
2.7 A cryo-EM structure of in vitro assembled type 1 pilus rod
Method: helical / : Hospenthal M, Zyla D, Glockshuber R, Waksman G

EMDB-17878: 
2.5 A cryo-EM structure of the in vitro FimD-catalyzed assembly of type 1 pilus rod
Method: helical / : Zyla D, Hospenthal M, Glockshuber R, Waksman G

PDB-8psv: 
2.7 A cryo-EM structure of in vitro assembled type 1 pilus rod
Method: helical / : Hospenthal M, Zyla D, Glockshuber R, Waksman G

PDB-8ptu: 
2.5 A cryo-EM structure of the in vitro FimD-catalyzed assembly of type 1 pilus rod
Method: helical / : Zyla D, Hospenthal M, Glockshuber R, Waksman G

EMDB-16813: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ
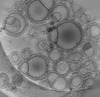
EMDB-16814: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ
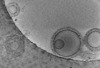
EMDB-16815: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ

EMDB-16794: 
Cryo-EM structure of the human GBP1 dimer bound to GDP-AlF3
Method: single particle / : Kuhm TI, Jakobi AJ

EMDB-18892: 
Lipid droplet-Vacuole contacts in Ldo16 overexpression yeast strain.
Method: electron tomography / : Collado J

EMDB-18893: 
Lipid droplet-vacuole and Nucleus-vacuole contacts in WT yeast cell starved for 4 hours
Method: electron tomography / : Collado J

EMDB-18894: 
Lipid droplet lipophagy in 4-hour starved WT yeast cell.
Method: electron tomography / : Collado J

EMDB-18895: 
Multiple vacuole-lipid droplet-nucleus contacts in 4-hour starved WT yeast cell.
Method: electron tomography / : Collado J

EMDB-18896: 
Lipophagy in 5-day starved WT yeast cell.
Method: electron tomography / : Collado J

EMDB-18897: 
Lipid droplets in proximity to a vacuole in dLdo strain cell after 5-day starvation.
Method: electron tomography / : Collado J

EMDB-18898: 
Vacuolar contents of WT cell after 5-day starvation.
Method: electron tomography / : Collado J

EMDB-18899: 
Lipid droplet-nucleus contacts in dLdo yeast strain after 5-day starvation.
Method: electron tomography / : Collado J

EMDB-17119: 
24-meric catalytic domain of dihydrolipoamide acetyltransferase (E2) of the E. coli pyruvate dehydrogenase complex.
Method: single particle / : Zdanowicz R, Meinhold S, Glockshuber R

EMDB-17126: 
E. coli pyruvate dehydrogenase (E1) in complex with dihydrolipoamide acetyltransferase (E2) peripheral subunit-binding domain.
Method: single particle / : Zdanowicz R, Meinhold S, Glockshuber R

PDB-8orb: 
24-meric catalytic domain of dihydrolipoamide acetyltransferase (E2) of the E. coli pyruvate dehydrogenase complex.
Method: single particle / : Zdanowicz R, Meinhold S, Glockshuber R

EMDB-17804: 
Bat-Hp-CoV Nsp1 and eIF1 bound to the human 40S small ribosomal subunit
Method: single particle / : Schubert K, Karousis ED, Ban I, Lapointe CP, Leibundgut M, Baeumlin E, Kummerant E, Scaiola A, Schoenhut T, Ziegelmueller J, Puglisi JD, Muehlemann O, Ban N

EMDB-17805: 
MERS-CoV Nsp1 bound to the human 43S pre-initiation complex
Method: single particle / : Schubert K, Karousis ED, Ban I, Lapointe CP, Leibundgut M, Baeumlin E, Kummerant E, Scaiola A, Schoenhut T, Ziegelmueller J, Puglisi JD, Muehlemann O, Ban N

PDB-8ppk: 
Bat-Hp-CoV Nsp1 and eIF1 bound to the human 40S small ribosomal subunit
Method: single particle / : Schubert K, Karousis ED, Ban I, Lapointe CP, Leibundgut M, Baeumlin E, Kummerant E, Scaiola A, Schoenhut T, Ziegelmueller J, Puglisi JD, Muehlemann O, Ban N

PDB-8ppl: 
MERS-CoV Nsp1 bound to the human 43S pre-initiation complex
Method: single particle / : Schubert K, Karousis ED, Ban I, Lapointe CP, Leibundgut M, Baeumlin E, Kummerant E, Scaiola A, Schoenhut T, Ziegelmueller J, Puglisi JD, Muehlemann O, Ban N

EMDB-33329: 
High resolution cry-EM structure of the human 80S ribosome from SNORD127+/+ Kasumi-1 cells
Method: single particle / : Cheng J, Beckmann R

EMDB-33330: 
High resolution cry-EM structure of the human 80S ribosome from SNORD127+/- Kasumi-1 cells
Method: single particle / : Cheng J, Beckmann R

PDB-7xnx: 
High resolution cry-EM structure of the human 80S ribosome from SNORD127+/+ Kasumi-1 cells
Method: single particle / : Cheng J, Beckmann R

PDB-7xny: 
High resolution cry-EM structure of the human 80S ribosome from SNORD127+/- Kasumi-1 cells
Method: single particle / : Cheng J, Beckmann R

EMDB-13404: 
Nanodisc reconstituted MsbA in complex with nanobodies, spin-labeled at position A60C
Method: single particle / : Januliene D, Parey K, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

EMDB-13405: 
AMP-PNP bound nanodisc reconstituted MsbA with nanobodies, spin-labeled at position A60C
Method: single particle / : Januliene D, Parey K, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

EMDB-13406: 
AMP-PNP bound nanodisc reconstituted MsbA with nanobodies, spin-labeled at position T68C
Method: single particle / : Januliene D, Parey K, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

EMDB-13409: 
Nanodisc reconstituted MsbA in complex with nanobodies, spin-labeled at position T68C
Method: single particle / : Januliene D, Parey K, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

PDB-7ph2: 
Nanodisc reconstituted MsbA in complex with nanobodies, spin-labeled at position A60C
Method: single particle / : Januliene D, Parey K, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

PDB-7ph3: 
AMP-PNP bound nanodisc reconstituted MsbA with nanobodies, spin-labeled at position A60C
Method: single particle / : Parey K, Januliene D, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

PDB-7ph4: 
AMP-PNP bound nanodisc reconstituted MsbA with nanobodies, spin-labeled at position T68C
Method: single particle / : Parey K, Januliene D, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

PDB-7ph7: 
Nanodisc reconstituted MsbA in complex with nanobodies, spin-labeled at position T68C
Method: single particle / : Parey K, Januliene D, Galazzo L, Meier G, Vecchis D, Striednig B, Hilbi H, Schaefer LV, Kuprov I, Bordignon E, Seeger MA, Moeller A

PDB-7tbl: 
Composite structure of the human nuclear pore complex (NPC) cytoplasmic face generated with a 12A cryo-ET map of the purified HeLa cell NPC
Method: subtomogram averaging / : Bley CJ, Nie S, Mobbs GW, Petrovic S, Gres AT, Liu X, Mukherjee S, Harvey S, Huber FM, Lin DH, Brown B, Tang AW, Rundlet EJ, Correia AR, Chen S, Regmi SG, Stevens TA, Jette CA, Dasso M, Patke A, Palazzo AF, Kossiakoff AA, Hoelz A

EMDB-14238: 
Cryo-EM structure of Bacillus megaterium gas vesicles
Method: helical / : Huber ST, Evers W, Jakobi AJ

EMDB-14340: 
Cryo-EM structure of Bacillus megaterium gas vesicles
Method: helical / : Huber ST, Evers W, Jakobi AJ

PDB-7r1c: 
Cryo-EM structure of Bacillus megaterium gas vesicles
Method: helical / : Huber ST, Evers W, Jakobi AJ

PDB-7tbm: 
Composite structure of the dilated human nuclear pore complex (NPC) generated with a 37A in situ cryo-ET map of CD4+ T cell NPC
Method: subtomogram averaging / : Bley CJ, Nie S, Mobbs GW, Petrovic S, Gres AT, Liu X, Mukherjee S, Harvey S, Huber FM, Lin DH, Brown B, Tang AW, Rundlet EJ, Correia AR, Chen S, Regmi SG, Stevens TA, Jette CA, Dasso M, Patke A, Palazzo AF, Kossiakoff AA, Hoelz A

EMDB-23949: 
The insulin receptor ectodomain in complex with a venom hybrid insulin analog - "head" region
Method: single particle / : Blakely AD, Xiong X, Kim JH, Menting J, Schafer IB, Schubert HL, Agrawal R, Gutmann T, Delaine C, Zhang Y, Artik GO, Merriman A, Eckert D, Lawrence MC, Coskun U, Fisher SJ, Forbes BE, Safavi-Hemami H, Hill CP, Chou DHC

EMDB-23950: 
The insulin receptor ectodomain in complex with four venom hybrid insulins - symmetric conformation
Method: single particle / : Blakely AD, Xiong X, Kim JH, Menting J, Schafer IB, Schubert HL, Agrawal R, Gutmann T, Delaine C, Zhang Y, Artik GO, Merriman A, Eckert D, Lawrence MC, Coskun U, Fisher SJ, Forbes BE, Safavi-Hemami H, Hill CP, Chou DHC

EMDB-23951: 
The insulin receptor ectodomain in complex with three venom hybrid insulin molecules - asymmetric conformation
Method: single particle / : Blakely AD, Xiong X, Kim JH, Menting J, Schafer IB, Schubert HL, Agrawal R, Gutmann T, Delaine C, Zhang Y, Artik GO, Merriman A, Eckert D, Lawrence MC, Coskun U, Fisher SJ, Forbes BE, Safavi-Hemami H, Hill CP, Chou DHC

PDB-7mqo: 
The insulin receptor ectodomain in complex with a venom hybrid insulin analog - "head" region
Method: single particle / : Blakely AD, Xiong X, Kim JH, Menting J, Schafer IB, Schubert HL, Agrawal R, Gutmann T, Delaine C, Zhang Y, Artik GO, Merriman A, Eckert D, Lawrence MC, Coskun U, Fisher SJ, Forbes BE, Safavi-Hemami H, Hill CP, Chou DHC

PDB-7mqr: 
The insulin receptor ectodomain in complex with four venom hybrid insulins - symmetric conformation
Method: single particle / : Blakely AD, Xiong X, Kim JH, Menting J, Schafer IB, Schubert HL, Agrawal R, Gutmann T, Delaine C, Zhang Y, Artik GO, Merriman A, Eckert D, Lawrence MC, Coskun U, Fisher SJ, Forbes BE, Safavi-Hemami H, Hill CP, Chou DHC

PDB-7mqs: 
The insulin receptor ectodomain in complex with three venom hybrid insulin molecules - asymmetric conformation
Method: single particle / : Blakely AD, Xiong X, Kim JH, Menting J, Schafer IB, Schubert HL, Agrawal R, Gutmann T, Delaine C, Zhang Y, Artik GO, Merriman A, Eckert D, Lawrence MC, Coskun U, Fisher SJ, Forbes BE, Safavi-Hemami H, Hill CP, Chou DHC

EMDB-12766: 
Single particle cryo-EM reconstruction of just the top two rings of a 40-mer assembly of recombinant yeast Hsp26 S207E mutant.
Method: single particle / : Muehlhofer M, Peters C, Kriehuber T, Kreuzeder M, Kazman P, Rodina N, Reif B, Haslbeck M, Weinkauf S, Buchner J

EMDB-12771: 
Single particle cryo-EM reconstruction of a 40-mer assembly of recombinant yeast Hsp26 S207E mutant.
Method: single particle / : Muehlhofer M, Peters C, Kriehuber T, Kreuzeder M, Kazman P, Rodina N, Reif B, Haslbeck M, Weinkauf S, Buchner J

EMDB-12772: 
Single particle cryo-EM reconstruction of a 40-mer assembly of recombinant yeast Hsp26 mutant S47ET48E.
Method: single particle / : Muehlhofer M, Peters C, Kriehuber T, Kreuzeder M, Kazman P, Rodina N, Reif B, Haslbeck M, Weinkauf S, Buchner J

EMDB-12773: 
Single particle cryo-EM reconstruction of a 40-mer assembly of recombinant yeast Hsp26.
Method: single particle / : Muehlhofer M, Peters C, Kriehuber T, Kreuzeder M, Kazman P, Rodina N, Reif B, Haslbeck M, Weinkauf S, Buchner J
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
